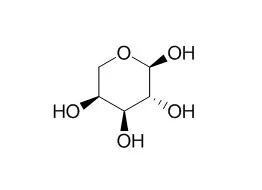
| Structure Identification: |
| FEBS J. 2008 Jun;275(11):2691-711. | | Biosynthesis of D-arabinose in mycobacteria - a novel bacterial pathway with implications for antimycobacterial therapy.[Pubmed: 18422659 ] |
METHODS AND RESULTS:
Decaprenyl-phospho-arabinose (beta-D-arabinofuranosyl-1-O-monophosphodecaprenol), the only known donor of D-Arabinose in bacteria, and its precursor, decaprenyl-phospho-ribose (beta-D-ribofuranosyl-1-O-monophosphodecaprenol), were first described in 1992. En route to D-arabinofuranose, the decaprenyl-phospho-ribose 2'-epimerase converts decaprenyl-phospho-ribose to decaprenyl-phospho-arabinose, which is a substrate for arabinosyltransferases in the synthesis of the cell-wall arabinogalactan and lipoarabinomannan polysaccharides of mycobacteria. The first step of the proposed decaprenyl-phospho-arabinose biosynthesis pathway in Mycobacterium tuberculosis and related actinobacteria is the formation of D-ribose 5-phosphate from sedoheptulose 7-phosphate, catalysed by the Rv1449 transketolase, and/or the isomerization of d-ribulose 5-phosphate, catalysed by the Rv2465 d-ribose 5-phosphate isomerase. d-Ribose 5-phosphate is a substrate for the Rv1017 phosphoribosyl pyrophosphate synthetase which forms 5-phosphoribosyl 1-pyrophosphate (PRPP). The activated 5-phosphoribofuranosyl residue of PRPP is transferred by the Rv3806 5-phosphoribosyltransferase to decaprenyl phosphate, thus forming 5'-phosphoribosyl-monophospho-decaprenol. The dephosphorylation of 5'-phosphoribosyl-monophospho-decaprenol to decaprenyl-phospho-ribose by the putative Rv3807 phospholipid phosphatase is the committed step of the pathway. A subsequent 2'-epimerization of decaprenyl-phospho-ribose by the heteromeric Rv3790/Rv3791 2'-epimerase leads to the formation of the decaprenyl-phospho-arabinose precursor for the synthesis of the cell-wall arabinans in Actinomycetales. The mycobacterial 2'-epimerase Rv3790 subunit is similar to the fungal D-arabinono-1,4-lactone oxidase, the last enzyme in the biosynthesis of D-erythroascorbic acid, thus pointing to an evolutionary link between the D-arabinofuranose- and L-ascorbic acid-related pathways. Decaprenyl-phospho-arabinose has been a lead compound for the chemical synthesis of substrates for mycobacterial arabinosyltransferases and of new inhibitors and potential antituberculosis drugs. The peculiar (omega,mono-E,octa-Z) configuration of decaprenol has yielded insights into lipid biosynthesis, and has led to the identification of the novel Z-polyprenyl diphosphate synthases of mycobacteria. Mass spectrometric methods were developed for the analysis of anomeric linkages and of dolichol phosphate-related lipids.
CONCLUSIONS:
In the field of immunology, the renaissance in mycobacterial polyisoprenoid research has led to the identification of mimetic mannosyl-beta-1-phosphomycoketides of pathogenic mycobacteria as potent lipid antigens presented by CD1c proteins to human T cells. |
|

 Cell. 2018 Jan 11;172(1-2):249-261.e12. doi: 10.1016/j.cell.2017.12.019.IF=36.216(2019)
Cell. 2018 Jan 11;172(1-2):249-261.e12. doi: 10.1016/j.cell.2017.12.019.IF=36.216(2019) Cell Metab. 2020 Mar 3;31(3):534-548.e5. doi: 10.1016/j.cmet.2020.01.002.IF=22.415(2019)
Cell Metab. 2020 Mar 3;31(3):534-548.e5. doi: 10.1016/j.cmet.2020.01.002.IF=22.415(2019) Mol Cell. 2017 Nov 16;68(4):673-685.e6. doi: 10.1016/j.molcel.2017.10.022.IF=14.548(2019)
Mol Cell. 2017 Nov 16;68(4):673-685.e6. doi: 10.1016/j.molcel.2017.10.022.IF=14.548(2019)